(Originally published at Planetary Technologies, 15 September, 2025.)
With lots of recent chatter about whether the energy transition is slowing down, or not, here is a brief discussion of a couple of relevant charts I haven't yet had time to write up properly. If the globe winds up installing only 10 TW of PV, we are reasonably far into the transition. But if we are on the way to 100 TW, then we are only just getting started, and the world will see installation rates, and thus manufacturing rates, that are as much as 10X higher than what we will have in 2025.
Last year in an installment of The Sun Has Won series (Research Note: Discerning Trends in PV Installation Data PDF), I published a comparison of global historical PV installation amounts with four different scenarios for future installation ranging from 10 TW to 100TW of total generating capacity. The purpose was to provide a means to quantitatively assess how far along we might be towards the equilibrium end state, i.e., the point at which we will have installed all the PV we are going to. Each of the scenarios assumes total PV installation follows a logistic curve, which enables an estimation of the maximum annual rate of installation. (See the PDF for specific methods and justifications, including comments on assuming a sigmoidal trajectory.) Historical installation data is sufficiently variable that is not possible to make an easy judgement about which scenario is most likely. That is,
1) Given variation in annual PV installation data, it is still not possible to distinguish which trajectory we are on.
2) Consequently, we cannot yet judge accurately what the endpoint will be, nor when it will arrive.
From the abstract of the Research Note:
There are at least three different historical price eras for PV, with different market dynamics, and there are today at least four distinct PV markets, with installation rates determined by contrasting local policy priorities. Due to this variability, different models of future market installation that are consistent with historical installation data can produce final installation totals, and maximum installation rates, that span more than an order of magnitude; existing data are consistent with a 2050 installation total of 25–100 TW, and a maximum installation rate of 600 GW to 6 TW per year. In other words, we cannot distinguish between a wide range of outcomes given existing data. Nevertheless, the exercise of comparing models to the historical record can help delineate and constrain the range of our ignorance, which provides a basis for evaluating scenarios for PV installation over the next three decades.
One point above needs further explanation before continuing. There is not a single global market for PV, but rather many different markets, with different installation bases, growth rates, and eventual equilibrium states. Thus lumping all the data into a single annual installation total, and looking at a single cumulative total, can only tell us so much. But it is still a useful exercise, because it informs us about the overall pace of change, which has very practical implications for annual demand and, therefore, for annual investment by, for example, manufacturers, customers, and utilities.
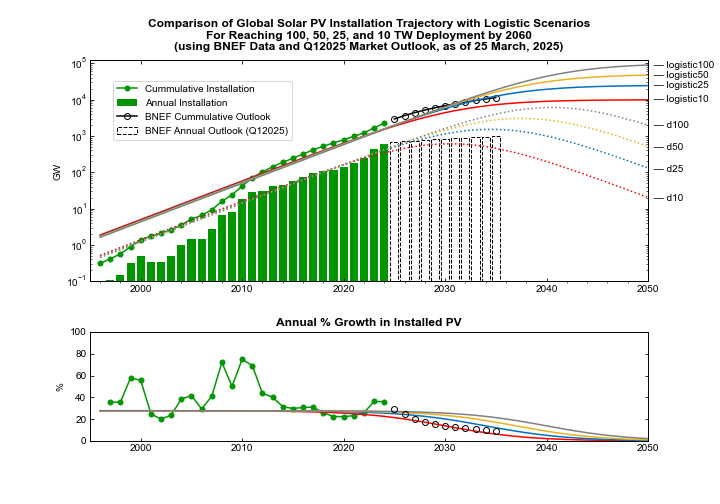
An important feature of this chart is the bottom panel, which compares the annual growth rate in installation to what you would expect for each of the four scenarios. The historical growth rate settled into a small range between 2014 and 2022, before jumping for the next two years. The stable range kept global installation on pace for anywhere between 10 TW and 100 TW, and the 2023 and 2024 rates ran substantially hotter than even the 100 TW pace.
The updated chart below compares the four scenarios with projections from the BloombergNEF (BNEF) Q1 2025 Market Outlook. Notably, BNEF projects a significant deceleration in the global PV installation rate starting next year. (I will just take this on its face value in what follows. Though I will note that for several years running the Market Outlook has forecast a sudden slowdown that has yet to materialize. But don’t judge that too harshly. Due to its customer base, BNEF has an organizational motivation to be conservative and err on the low side. As always, hats off to the BNEF team for gathering — and publishing — the best data set out there.)
Top panel: Four scenarios for future global PV installation, ranging from 10 TW to 100 TW. Historical data for annual installation (green bars) and cumulative PV capacity (solid green line through data points); the four scenarios for future annual PV installation (dotted lines) and future cumulative capacity (solid lines) are each consistent with historical data. Additionally, the installation forecast from the BNEF Q1 2025 Market Outlook is shown as open boxes with dashed black outline, and the resulting cumulative forecast shown as open black circles. The scenarios are not quantitatively fit to the historical data. Bottom panel: The annual percentage increase in installed PV for the same four scenarios plotted in comparison to historical percentage increases. The percentage increase implicit in the BNEF Q1 202t installation forecast out to 2035 is shown as open black circles (Source: BloombergNEF, Planetary Technologies).
The primary consequence of a sudden deceleration in PV installation would be hewing closer to a much lower final installation total. Similarly, the global annual market for PV would not expand as fast as it has been, and there would be less need for investment in manufacturing capacity. Yet it is unclear, given the economic advantage of PV over all other electricity generation technologies, why installation should decelerate. As I conclude in the aforementioned Research Note:
Given the economic advantage of solar over all other generation sources, particularly when coupled to batteries, it would be more surprising for solar installations to slow down than it would be for solar installations to continue surpassing expectations.
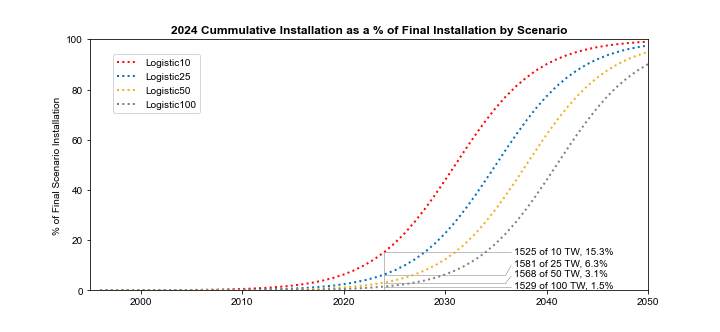
Finally, to try to put the four scenarios in historical context, here is a previously unpublished chart that estimates how far we are along each scenario based on historical data. If we are destined to install only 10 TW, then we are already about 15% of the way there, and a transition to linear growth should happen soon. But if we going to see 100 TW of solar installed by between 2050 and 2060, then we are only 1.5% into the transition, and we should expect to see at least another ten years of exponential growth.
Progress into the energy transmission depends on the end state. As of the end of 2024, we would be ~15% towards an equilibrium state with 10 TW of PV installed, but only ~1.5% towards an equilibrium state with 100 TW installed (Source: Planetary Technologies).
I will finish for now with another quote from the Research Note, which stands as a decent summary of the state of what we can say about how fast we are going, and where we are going, at least until we have several more years of data.
The 10 TW scenario is characterized by a maximum installation rate of 620 GW per year, occurring in 2030, and the 100 TW scenario is characterized by a maximum installation rate of 6 TW per year, in 2040. Notably, by the end of 2022 there was already nearly 1 TW of global manufacturing capacity for modules, although in the first half of 2024 some factories are running at only 40% capacity due to low prices. I estimate that global annual manufacturing capacity today is about 1.5 TW. China by itself is forecast to have 1.7 TW of annual manufacturing capacity by 2026. Given existing and future manufacturing capacity, and given the current volume of warehoused panels, near-term installation rates will not be constrained by supply but rather by other factors, such as installation labor availability, policy, and transmission capacity. Assuming that the real-world installation trajectory will be described by a logistic curve, achieving a 1 TW annual installation rate by 2030 is broadly consistent with a total final installation of between 25 and 100 TW. In the context of aggressive installation policies around the world, with prices unlikely to rise much due to oversupply, it would be bold to predict that installation rates will fall in the immediate future.